Send a Brain. Get the Answers.

Mouse Brain - Allen Institute
Understanding brain circuits is still painfully slow.
Modern neuroscience can record activity, sequence genes, and manipulate neurons with precision. But measuring how circuits are physically wired remains extremely difficult. Getting this data requires months of work and massive computational reconstruction efforts.
For example, mapping the FlyWire connectome, the complete wiring diagram of a fruit fly brain (see video) with just 140,000 neurons, took over 5 years and required more than 33 person-years of proofreading effort from dozens of researchers worldwide, plus millions of hours of computation.
As a result, many studies rely on indirect measurements instead of directly quantifying circuit structure simply because the direct approach is too expensive and time-consuming.
FlyWire - Animation by Tyler Sloan
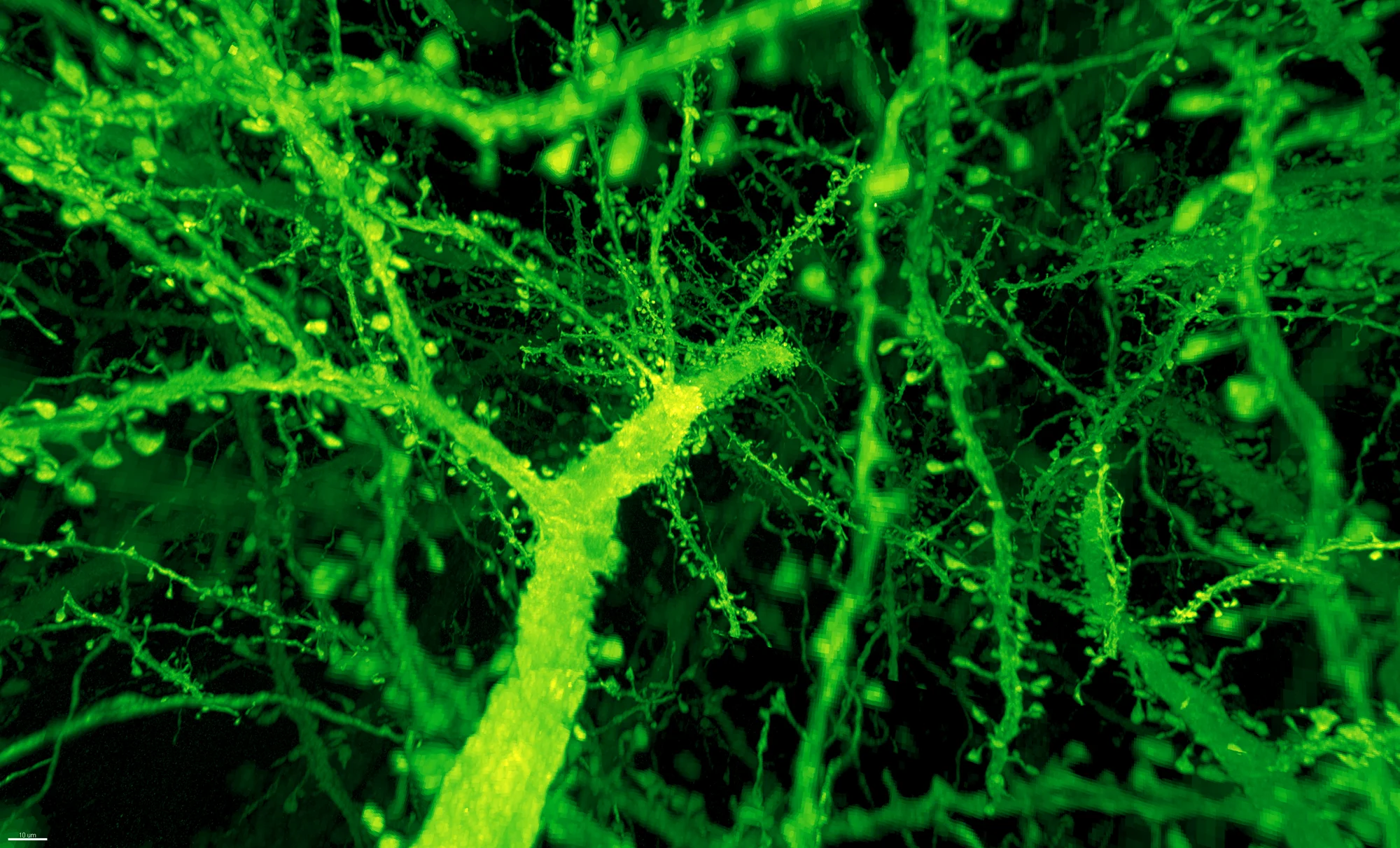
Gao et al. / Science 2019
But what if it wasn't?
The reason FlyWire took so much time and effort was because it required electron microscopy, a painstaking process involving physical sectioning, ultra-high-resolution imaging, and massive computational reconstruction of individual neurons that still needed manual proofreading. This approach delivers unparalleled detail but is prohibitively slow and expensive for most research questions.
Expansion-assisted light sheet fluorescence microscopy offers a fundamentally different approach: physically expanding the tissue and using fluorescent markers to image synapses directly, bypassing the need for electron microscopy entirely.
This makes circuit mapping orders of magnitude faster and more accessible. But raw images alone aren't enough, they need to be transformed into actionable structural data.
That's exactly what our pipeline strives to achieve.
The Pipeline
From tissue to data — a streamlined approach to neural circuit mapping

Tissue preparation
Brain tissue is physically expanded and fluorescently labeled to reveal synaptic structures.

Large-scale imaging
Light sheet microscopy rapidly captures entire neural circuits across expanded brain volumes.

Computational reconstruction
Machine learning algorithms detect and map synapses throughout the imaged tissue.

Circuit-level outputs
Researchers receive quantitative metrics describing how circuits are organized at synapse resolution.
Brain tissue is physically expanded and fluorescently labeled to reveal synaptic structures.
Light sheet microscopy rapidly captures entire neural circuits across expanded brain volumes.
Machine learning algorithms detect and map synapses throughout the imaged tissue.
Researchers receive quantitative metrics describing how circuits are organized at synapse resolution.
Road to the Proof of Concept
Goal: End-to-end validation of the pipeline: expansion, imaging and data processing on a Drosophila brain.
Let's build this together
We're looking for people who get excited about ambitious technical challenges and want to help create something meaningful from day one.
WHERE YOU MIGHT FIT IN
Neuroscience & Biology
Tissue processing, expansion microscopy, fluorescent labeling — the wet-lab backbone of the pipeline.
Optics & Imaging
Light sheet microscopy, optical design, and large-scale volumetric acquisition.
Computer Science & ML
Image processing, 3D reconstruction, synapse detection, and building the software that ties it all together.
Engineering & Hardware
Custom instrumentation, automation, and scaling the physical setup.
You don't need to have experience in every area. What matters most is curiosity, drive, and a willingness to learn and experiment. This is early-stage work — we're figuring things out together, learning as we go.
Based at DTU (Technical University of Denmark)
Let's build this together
We're looking for people who get excited about ambitious technical challenges and want to help create something meaningful from day one.
You don't need to have experience in every area. What matters most is curiosity, drive, and a willingness to learn and experiment. This is early-stage work — we're figuring things out together, learning as we go.
Based at DTU (Technical University of Denmark)
WHERE YOU MIGHT FIT IN
Neuroscience & Biology
Tissue processing, expansion microscopy, fluorescent labeling — the wet-lab backbone of the pipeline.
Optics & Imaging
Light sheet microscopy, optical design, and large-scale volumetric acquisition.
Computer Science & ML
Image processing, 3D reconstruction, synapse detection, and building the software that ties it all together.
Engineering & Hardware
Custom instrumentation, automation, and scaling the physical setup.